// Selected work
Agents, pipelines, and research projects
Five projects that cover the range: open-source agentic AI, agent-callable genomics tools, production multi-agent systems, self-optimising ML pipelines, and applied research. Each links to code, a demo, or a writeup where possible.
Flagship · Open source
LangGraph · Claude · Python
agentic-genomics · GenomicsCopilot
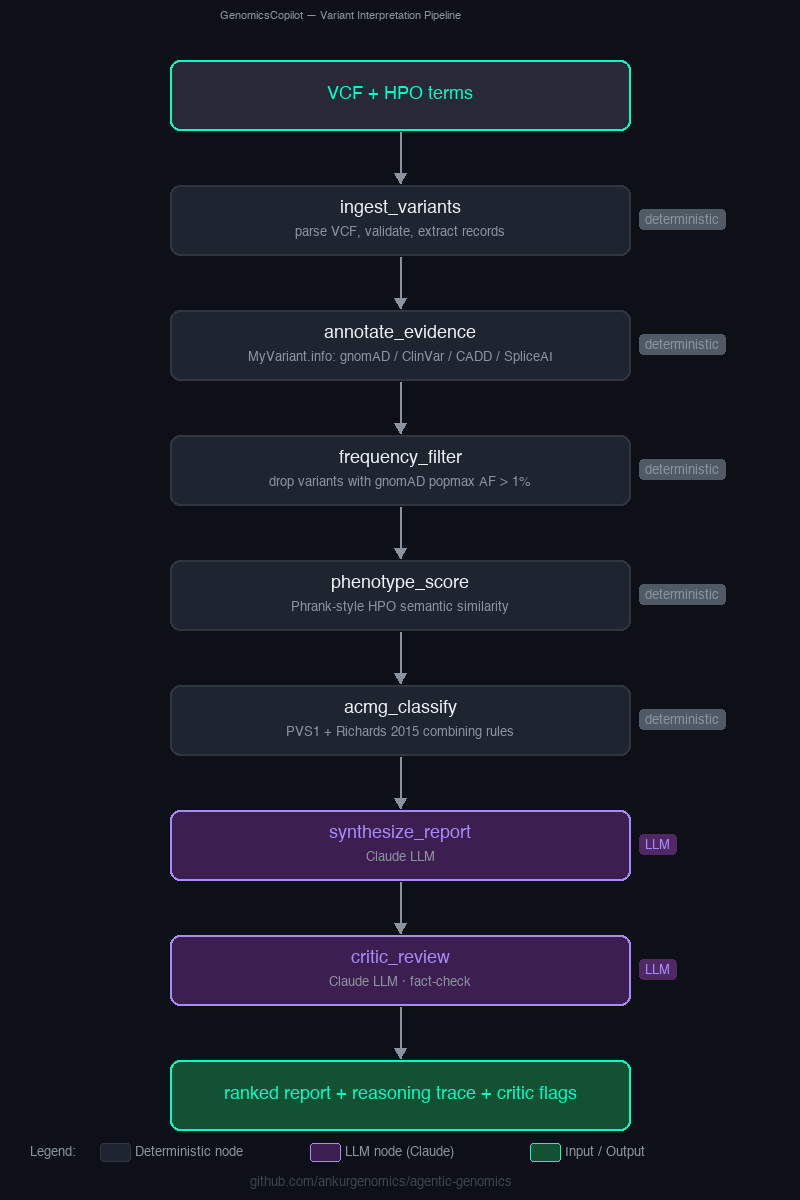
An open-source LangGraph agent for explainable variant interpretation — every call leaves a full reasoning trace a human can audit. Research demonstration, not clinical.
Takes a VCF + HPO phenotype terms and returns a ranked, explainable report of candidate variants. Deterministic nodes handle ingest, gnomAD / ClinVar / SpliceAI lookups via MyVariant.info, a transparent ACMG-lite rule engine (7 criteria, a proper PVS1 check, and Richards-et-al-2015 combining rules), and a Phrank-style HPO semantic-similarity score. An LLM synthesiser ranks candidates and writes the narrative; a second LLM critic fact-checks those claims against the evidence JSON and flags anything unsupported. Every run emits a machine-readable reasoning trace. See LIMITATIONS.md for an honest accounting of what this system does not do.
7 nodes
LangGraph + critic review
4 tools
MyVariant · Phrank HPO · ACMG-lite · critic
MIT
Open source, Python 3.11+
LangGraph
Claude / Anthropic
Pydantic v2
pysam
Streamlit
Typer CLI
GitHub Actions
Open source · Agent skills
Python · Claude Haiku · REST APIs
genomics-skills — Agent-Callable Skill Library
8 pure-Python genomics skills that downstream agents can call: expression profiling, survival analysis, protein mapping, pathway enrichment, literature search, and more.
The downstream skill layer for agentic-genomics. Each skill is a standalone, agent-discoverable Python module with a SKILL.md contract, CLI entrypoint, and deterministic output (TSV + PNG/SVG). Pan-cancer expression uses real TCGA data (9,479 samples across 31 cancer types via cBioPortal). Kaplan-Meier survival runs Cox PH regression on actual patient data. LLM-powered routing via Claude Haiku maps natural-language queries to the right skill. Parquet caching makes repeat queries instant.
8 skills
Agent-callable, SKILL.md contract each
9,479
Real TCGA patient samples
MIT
Open source, Python 3.9+
Python
Claude Haiku
cBioPortal API
MyVariant.info
NCBI E-utils
PDB / AlphaFold
Pandas
Matplotlib
GenomicsOps AI
Five specialized agents that triage and resolve DRAGEN, ICA and SGE/HPC pipeline failures end-to-end.
Personal project built on weekends: Trigger → Log Fetcher → RAG → Classifier → JIRA Writer. Tested on real failure scenarios — BED-file overlaps, samplesheet index mismatches, stuck SGE jobs — with end-to-end triage and ticket creation.
Multi-agent
Claude API
RAG
Python
JIRA & Confluence APIs
Side project · happy to walk through architecture in interviews
Production · Cloud
Autonomous Genomic Pipelines (Mirxes)
Self-optimising WGS/RNA-seq workflows on AWS with adaptive resource allocation and automated QC gating.
Designed and shipped the AWS infrastructure for the Singapore National Precision Medicine project. Nextflow on AWS Batch, with Lambda + Step Functions orchestrating sample intake, QC decisions, and output delivery. Processed 6,000+ samples with minimal human intervention.
400 TB
Genomic data managed
Nextflow
AWS Batch
Lambda
Step Functions
Docker
IaC
Research · PhD
Age-dependent hepatocyte epigenomics
Integrative RNA-seq + ChIP-seq + Hi-C analysis pipeline revealing age-driven chromatin reorganisation in mouse liver.
PhD work at NTU: built end-to-end NGS analysis pipelines for transcriptome, histone modifications, and 3D chromatin. Identified H3K27me3 as a key age-dependent regulator. The technical stack — reproducible pipelines, multi-omic integration, careful statistics — is the same foundation I now use for agentic ML systems.
RNA-seq
ChIP-seq
Hi-C (3C-seq)
R · Bioconductor
Python
k-means / GSEA
