// Selected work
Agents, pipelines, and research projects
Six projects that cover the range: open-source agentic AI for outbreak surveillance, agent-callable genomics tools, production multi-agent systems, self-optimising ML pipelines, and applied research. Each links to code, a demo, or a writeup where possible.
Flagship · Open source
LangGraph · Claude · Python
agentic-genomics · GenomicsCopilot
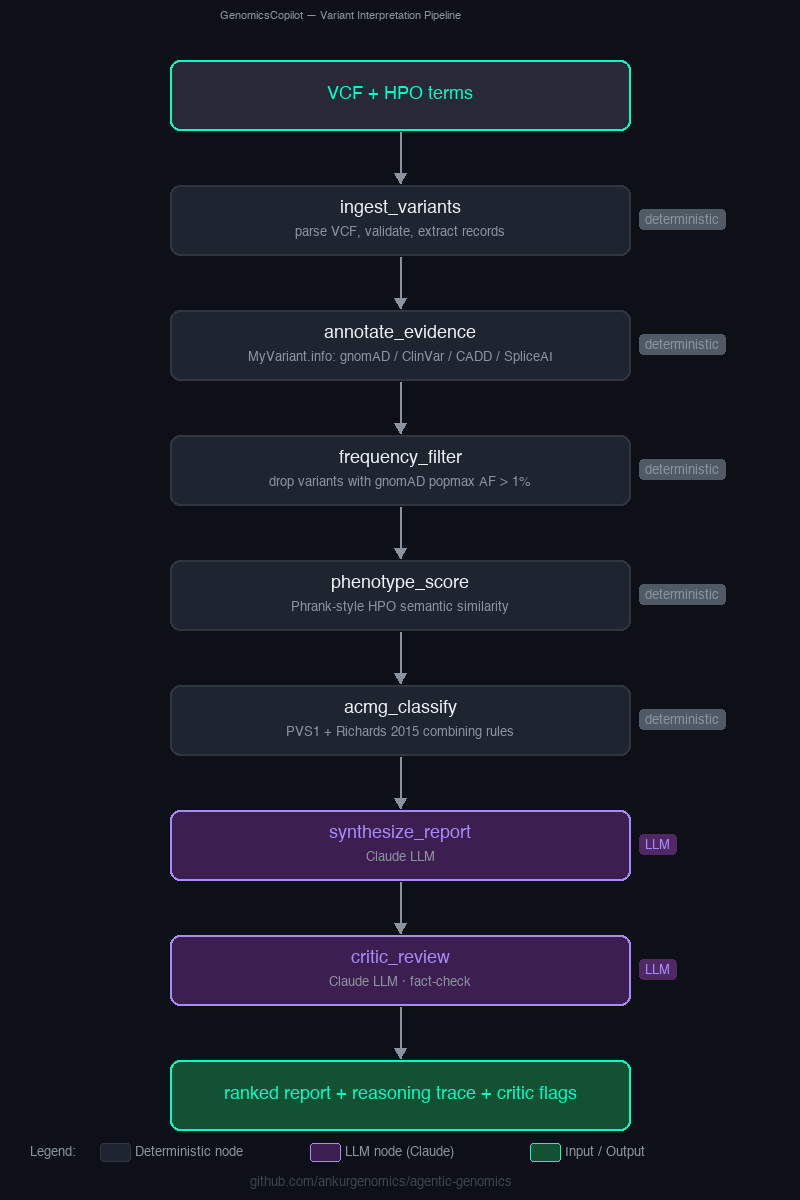
An open-source LangGraph agent for explainable variant interpretation — every call leaves a full reasoning trace a human can audit. Research demonstration, not clinical.
Takes a VCF + HPO phenotype terms and returns a ranked, explainable report of candidate variants. Deterministic nodes handle ingest, gnomAD / ClinVar / SpliceAI lookups via MyVariant.info, a transparent ACMG-lite rule engine (7 criteria, a proper PVS1 check, and Richards-et-al-2015 combining rules), and a Phrank-style HPO semantic-similarity score. An LLM synthesiser ranks candidates and writes the narrative; a second LLM critic fact-checks those claims against the evidence JSON and flags anything unsupported. Every run emits a machine-readable reasoning trace. See LIMITATIONS.md for an honest accounting of what this system does not do.
7 nodes
LangGraph + critic review
4 tools
MyVariant · Phrank HPO · ACMG-lite · critic
MIT
Open source, Python 3.11+
LangGraph
Claude / Anthropic
Pydantic v2
pysam
Streamlit
Typer CLI
GitHub Actions
Open source · Agentic AI
LangGraph · Python · No API key
outbreak-agent -- Infectious Disease Triage Pipeline
4-node LangGraph state machine that triages infectious disease cases -- built around the April 2026 MV Hondius / Andes virus event, the first confirmed human-to-human hantavirus transmission on a cruise ship.
A self-correcting agentic pipeline: genomic_node identifies clade and mutations, linkage_node resolves contact clusters and transmission mode, risk_node scores 0-100 with tier (LOW / MEDIUM / HIGH / CRITICAL), and critic_node enforces four consistency rules -- looping back up to 3 times if flags are raised. Generates a 3-panel matplotlib risk dashboard (PNG) and a structured A4 PDF triage report automatically on every run. No API key. No cost. 33 tests, fully deterministic. Part of the same ecosystem as gwas_nf: gwas_bridge.py feeds REGENIE tophits directly into this triage pipeline for population-to-clinical interpretation.
4 nodes
genomic / linkage / risk / critic
33 tests
23 unit + 10 integration, free to run
<2 sec
CRITICAL triage, MV Hondius case
LangGraph 0.6
LangChain
matplotlib
ReportLab
pytest
Apache 2.0
Open source · Agent skills
Python · Claude Haiku · REST APIs
genomics-skills — Agent-Callable Skill Library
8 pure-Python genomics skills that downstream agents can call: expression profiling, survival analysis, protein mapping, pathway enrichment, literature search, and more.
The downstream skill layer for agentic-genomics. Each skill is a standalone, agent-discoverable Python module with a SKILL.md contract, CLI entrypoint, and deterministic output (TSV + PNG/SVG). Pan-cancer expression uses real TCGA data (9,479 samples across 31 cancer types via cBioPortal). Kaplan-Meier survival runs Cox PH regression on actual patient data. LLM-powered routing via Claude Haiku maps natural-language queries to the right skill. Parquet caching makes repeat queries instant.
8 skills
Agent-callable, SKILL.md contract each
9,479
Real TCGA patient samples
MIT
Open source, Python 3.9+
Python
Claude Haiku
cBioPortal API
MyVariant.info
NCBI E-utils
PDB / AlphaFold
Pandas
Matplotlib
GenomicsOps AI
Five specialized agents that triage and resolve DRAGEN, ICA and SGE/HPC pipeline failures end-to-end.
Personal project built on weekends: Trigger → Log Fetcher → RAG → Classifier → JIRA Writer. Tested on real failure scenarios — BED-file overlaps, samplesheet index mismatches, stuck SGE jobs — with end-to-end triage and ticket creation.
Multi-agent
Claude API
RAG
Python
JIRA & Confluence APIs
Side project · happy to walk through architecture in interviews
Open source · Production · Cloud
Nextflow · REGENIE · AWS
gwas_nf — Multi-Ethnic GWAS Pipeline
REGENIE-based Nextflow GWAS pipeline for the TEMUS multi-ethnic cohort (4 groups, 10,000 samples, 100,000 variants, 13 phenotypes) -- with downstream agentic interpretation via GenomicsCopilot.
End-to-end population genomics pipeline: genotype QC and LD pruning, REGENIE whole-genome regression (Step 1 + Step 2), multi-ethnic stratified association, interactive Manhattan plots per phenotype per group, and automated HTML reports. Top hits feed into GenomicsCopilot via gwas_bridge.py: variant IDs extracted from .regenie.filtered.gz tophit files and passed to the 7-node LangGraph interpretation pipeline for ACMG classification and LLM-powered clinical synthesis. Population discovery to clinical interpretation in a single command.
4 groups
TEMUS multi-ethnic cohort
13 phenotypes
Parallel GWAS runs
100k variants
REGENIE whole-genome regression
Nextflow DSL2
REGENIE
AWS Batch
Python
R / ggplot2
Docker
Research · PhD
Age-dependent hepatocyte epigenomics
Integrative RNA-seq + ChIP-seq + Hi-C analysis pipeline revealing age-driven chromatin reorganisation in mouse liver.
PhD work at NTU: built end-to-end NGS analysis pipelines for transcriptome, histone modifications, and 3D chromatin. Identified H3K27me3 as a key age-dependent regulator. The technical stack — reproducible pipelines, multi-omic integration, careful statistics — is the same foundation I now use for agentic ML systems.
RNA-seq
ChIP-seq
Hi-C (3C-seq)
R · Bioconductor
Python
k-means / GSEA
